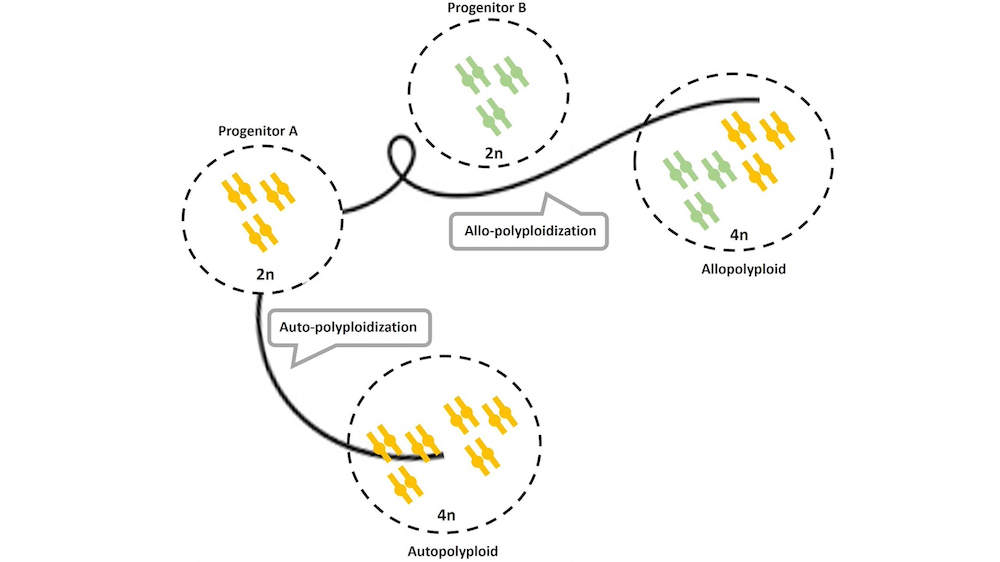
Polyploidization is a recurrent evolutionary phenomenon that generates diversity. Accidental polyploidization events are one of the initial steps before adaptation during periods of environmental change. Many plants, animals, fungi and other eukaryotes emerged due to ancient or recent polyploidization events. The effects of polyploidization in the genome, proteome, metabolome and its relationship with the new adaptive phenotype, are not well understood and multidisciplinary approaches are required.
In this Symposium, we bring experts in different disciplinary areas that might help us to unravel the effects of polyploidization.
Our speakers will be:
- David Peris, IATA-CSIC and University of Oslo. Expert in yeast comparative genomics. Organizer and project leader of the RCN PloidYeast, which sponsors this Symposium.
- Filip Kolář, Charles University in Prague. Expert in polyploidization in plants.
- Rike Stelkens, Stockholm University. Expert in experimental evolution, hybridization and fitness effects.
- Ana Conesa, I2SysBio-CSIC. Expert in multiomic integration.
- Héctor García, Staff Scientist (Quantitative Metabolic Modeling) Lawrence Berkeley National Laboratory (LBNL); Director (Data Science and Modeling) Joint BioEnergy Institute (JBEI); External Scientific Member Basque Center for Applied Mathematics (BCAM). Expert in Machine learning, synthetic biology and automation.
Program
09:00 Registration
09:30 Introduction Symposium by David Peris
09:45 Polyploidization by Filip Kolář
10:30 Hybridization by Rike Stelkens
11:15 Coffee break
11:45 Integromics by Ana Conesa (virtual)
12:30 Machine learning by Héctor García
The event is open to everyone interested in the topic until Arne Næs Auditorium 103 is filled, maximum 150 people. Opportunities for asking questions will be available.
The Symposium is supported by the UiO:Life Science and UiO:Energy and Environment