In this bioinformatics project, the student will use network analysis to analyze the 3D genome structures derived from various human tissues. The student will first use computational tools to calculate different centrality measures of the nodes in the network, including network degree, betweenness centrality, and closeness centrality. The student will associate these centrality measures with gene expression, to get an estimate of whether certain locations in the 3D genome structure lead to increase/decrease in gene expression levels. Finally, the student will perform network topology analysis, identifying modules of genome domains that are connected together and that are likely to act together in regulating gene expression.
Bioinformatics: Network analysis of the 3D genome
Chromosomes are organized in three dimensions (3D) inside the cell nucleus. This organization is crucial to maintain normal functions in the cell. Recently, new technologies allow high-throughput mapping of DNA contact points across the entire genome. Since this information sheds light on how parts of the DNA are interconnected, we can represent these models as large networks, and use tools from network analysis to better understand the topological features in the 3D genome. This knowledge is important for understanding when (and how) changes to these topologies can cause diseases such as cancer.
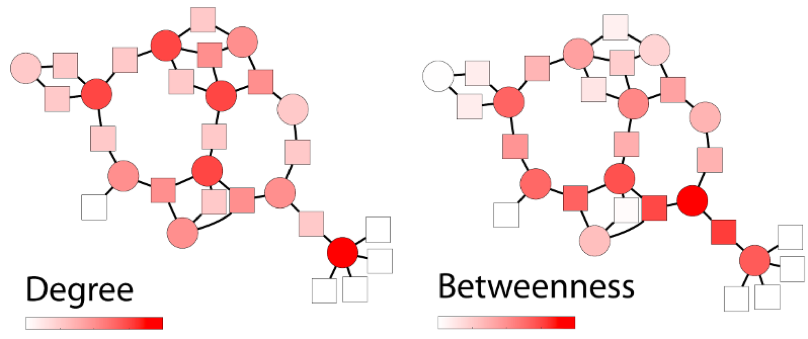
Figure 1: Values of different centrality measures in an example network, ranging from low (white) to high (red).
Publisert 28. sep. 2021 10:03
- Sist endret 28. sep. 2021 10:03